Provided at OPENSCREEN-GR by
- Agricultural University of Athens
- Biomedical Research Foundation, Academy of Athens
Services
Bioinformatic Analysis
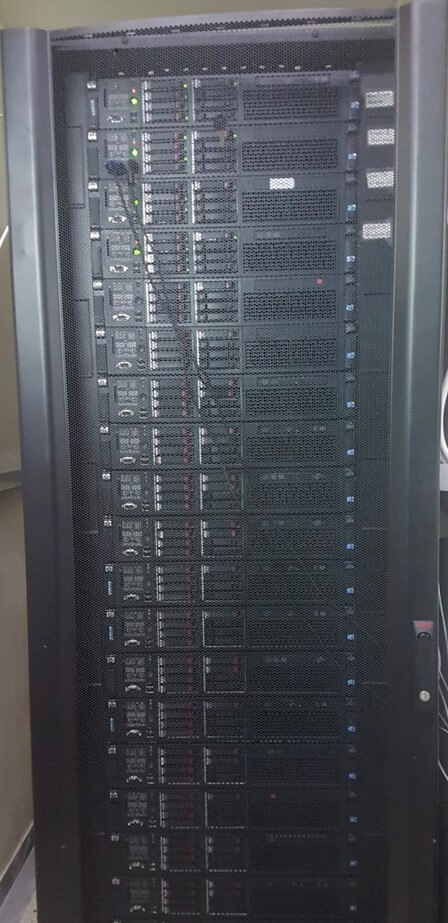
Super-Computer, Agricultural University of Athens
Equipment
For the purposes of developing the processing center for the bio-accounting analysis and provision of internet services, the host super-computer has been installed in a specially designed space. The server supercomputer architecture is based on multitask operations protocols.
Characteristics
- The server consists of 24 interconnected 16-core Intel Xeon 2.8GHz PCs with 32GB of internal memory each to manage search and programming tasks and databases.
Software
The following software and programming languages have been installed for the purpose of serving users in person and online:
- Windows 10 Windows Server 2016 S2
- Ubuntu 14.04 LTS
- Ubuntu 16.04 LTS
- Ubuntu 18.04 LTS
- XAMPP
- PHP Programing Language
- Java Script Programming Language
- HTML5 – Web Programming Language
- Python
- MySQL Programing Language
- MariaDB Programming Language
- MATLAB
Applications
The following computer applications have been installed for the purposes of the project:
- Molecular Operating Environment (MOE)
- LigandScout
- JalView
- Bambe-V2.01beta
- Pymol ™ v.2.0.6
- MolView
- Visual Studio 2015
- WinCoot v.0.8.9.2 • Phenix v.1.19.1
- Chimera v.1.15
- Shelxle v.1215
- Olex2 v.1.3.0
- Mercury-2020.3.0
- Tanimoto

Computational Techniques
Beis Lab - Biomedical Research Foundation, Academy of Athens
The Beis laboratory uses the zebrafish (Danio rerio), a small tropical fish, as an animal model to create experimental models of human disease. The zebrafish provides a wealth of benefits for Biomedical Research and in particular the ability to study signaling pathways at the organism level in vivo.
Drug Design and Molecular Modeling Lab - Biomedical Research Foundation, Academy of Athens
Drug Design and Molecular Modeling lab (Cournia lab) uses computational techniques to design new drug candidates. Specifically, techniques for computationally predicting the structure of a ligand-protein complex, for predicting the free binding energy of a drug candidate to a protein, de novo designing new drug candidates, exploring the dynamics of proteins that are pharmacological targets, properties of chemical molecules, and the construction of chemotherapeutic agents.
Database
Hippocrates Database, Agricultural University of Athens
The website of the sub-project was created at the address http://www.openscreen.aua.gr/. The portal cooperates with the specialized database of chemical compounds through advanced internet protocols and password encryption system. In the final stage of the development of the web portal, intelligent algorithms were applied for the display of data in the various fields where a large volume of information had been gathered, as well as for its parallel use by different users in separate terminals.
Finally, a specialized protocol was developed that includes various parameters for the continuous updating and development of both the web portal and the application database.
Application
From this website, authorized users can go and use the Hippocrates database
